As the world is more than half a year into the COVID-19 pandemic, doctors and researchers have a fairly good idea of what the main symptoms of the disease look like: cough, fever, shortness of breath, and fatigue, among others. But equally important to treating symptoms is understanding what the coronavirus that causes COVID-19, SARS-CoV-2, is doing inside human cells to make people so sick.
Like all viruses, SARS-CoV-2 breaks into a cell and hijacks its resources and machinery to create more viruses. Evolutionarily speaking, successful viruses are those that can effectively evade a cell’s defenses, but refrain from killing the cell outright (after all, the virus needs the cell to remain alive to be able to reproduce).
Human cells (and, more broadly, mammalian cells) have built-in defense mechanisms to deal with viral infections. The presence of viral genetic material in a cell triggers a cascade of events that lead to the production and secretion of a group of proteins called interferon, which will try to shut down the infection and notify neighboring cells of the threat. Researchers have found that patients with severe COVID-19 symptoms also show low levels of interferon response, suggesting that the interferon response is crucial for combatting the virus. How does the virus suppress these normal defense mechanisms?
A team led by Caltech researchers has now pinpointed the mechanisms through which the SARS-CoV-2 virus incapacitates human cells, essentially disabling the cell’s alarm system so that it cannot call for help or warn nearby cells of the infection. Understanding how the virus causes dysfunction at the cellular level gives new insights into how to fight it.
The research was conducted primarily in the laboratory of Mitchell Guttman, professor of biology and Heritage Medical Research Institute investigator. A paper describing the research appears online ahead of publication in the journal Cell.
The SARS-CoV-2 virus produces about 30 viral proteins. In this new research, the Guttman laboratory examined each of these and mapped out how they interact with the molecular components within human cells grown in a lab dish. They found that SARS-CoV-2 proteins attack three critical cellular processes to disrupt human protein production.
“Viruses are amazing,” says Emily Bruce, faculty scientist at the University of Vermont and a co-first author on the paper. “Viruses and host cells are continually in an evolutionary arms race to outwit one another. SARS-CoV-2 has evolved intricate and specific ways to disable cells without killing them outright, so that the virus can still use the cell for its own purposes.”
Some basic cell biology background first: The cell’s nucleus houses its genetic material, written as DNA. This so-called genome can be thought of as a comprehensive instruction manual, with “chapters” that might be titled “How to Send a Signal” or “What to Do in Case of Viral Infection,” for example. The rest of the cell contains the machinery that creates the proteins (such as interferon) that carry out these instructions.
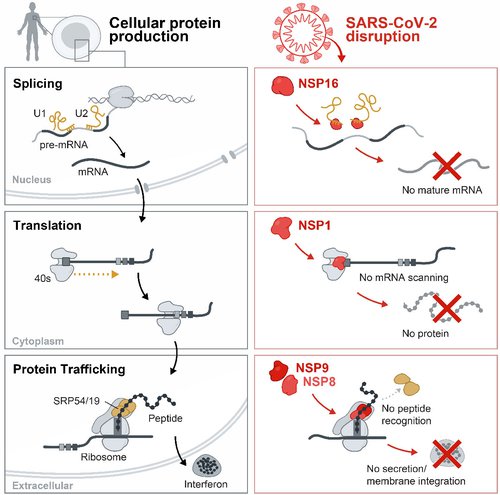
Credit: Inna-Marie Strazhnik / Caltech
The process for turning DNA instructions into useful proteins is called the “central dogma” of biology. The first step is transcription, through which a piece of DNA in the cell’s nucleus is read and copied into a form (a molecule called mRNA) that can leave the nucleus and travel to the rest of the cell. Before export out of the nucleus, mRNA is often re-assembled and “matured” in a process called splicing (top row).
After the mRNA is exported out of the nucleus, a piece of cellular machinery called the ribosome attaches to the mature mRNA, reads it, and builds the corresponding protein through a process called translation (middle row).
Some of these proteins are designed to move outside the cell of origin to transmit messages to other cells, for example, to warn about the presence of a viral infection. In this situation, another piece of cellular machinery called the signal recognition particle comes into play; it works as a kind of transport system that helps proteins move from inside to outside of a cell. This is known as protein trafficking (bottom row).
The Guttman lab discovered that SARS-CoV-2 proteins interfere with this whole process at multiple stages. Some of the virus’s proteins prevent mRNA from being fully spliced and properly assembled. Others plug up the ribosome so that it cannot form new proteins. Still other SARS-CoV-2 proteins interfere with the signal recognition particle and block protein transport.
The protein that plugs up the ribosome is called NSP1. Remarkably, the team found, NSP1 blocks human mRNA from entering the ribosome, but allows viral mRNA to pass through just fine. Viral mRNA contains a genetic signature at the beginning of each of its mRNAs that acts like an access code that effectively hijacks the ribosome to make viral proteins but not human proteins. Because viral production depends on this signature, it could represent a potent target for anti-viral therapeutic development.
“Each of the processes that SARS-CoV-2 disrupts— splicing, translation, and protein trafficking—is so important for converting the human genetic material into proteins, and they are essential for human biology,” says Guttman. “In fact, discovery of each of these processes has separately led to the awarding of a Nobel Prize. These are machines that are central to life. We cannot exist without them. SARS-CoV-2 has evolved in very specific ways to disable these cellular machines and disrupt their functions.”
“Our study illustrates the importance of basic science research, and establishes a pipeline to address newly emerging RNA viruses in the future,” says co-first author Abhik Banerjee, a graduate student in the Guttman laboratory. “Additionally, it illustrates the collaborative atmosphere of science at Caltech and elsewhere in the scientific community at its best. Here at Caltech, we have access to leaders in several keystone areas of biology, including professors Rebecca Voorhees (co-author on the published manuscript), Bil Clemons, and Shu-ou Shan in structural biology, all of whom were willing to discuss ramifications of our data and provide expertise in this relatively new area for us.”
Mario Blanco, a research scientist in the Guttman laboratory, agrees.
“Our ability to interrogate the human RNA targets of SARS-CoV-2 proteins allowed us to identify these mechanisms without prior evidence,” he says. “The methods and practices we developed here will allow us to apply these same processes to emergent diseases and even currently existing viruses where we lack a deep understanding of mechanism.”
The paper is titled “SARS-CoV-2 disrupts splicing, translation, and protein trafficking to suppress host defenses.”
Graduate student Abhik Banerjee, research scientist Mario Blanco, and faculty scientist Emily Bruce are the study’s co-first authors. Graduate student Drew Honson, graduate student Linlin Chen, and senior research scientist Amy Chow are co-second authors. Additional co-authors are graduate student Prashant Bhat, postdoctoral scholar Noah Ollikainen, former Guttman laboratory graduate student Sofia Quinodoz (PhD ’20), Colin Loney of the MRC-University of Glasgow Centre for Virus Research, Caltech research technician assistant Jasmine Thai, Zachary Miller of the University of Vermont College of Medicine, Aaron Lin of the Broad Institute of MIT and Harvard, Madaline Schmidt of the University of Vermont, Douglas Stewart of the MRC-University of Glasgow Centre for Virus Research, Daniel Goldfarb of the MRC-University of Glasgow Centre for Virus Research, Giuditta De Lorenzo of the MRC-University of Glasgow Centre for Virus Research, Suzannah Rihn of the MRC-University of Glasgow Centre for Virus Research, Caltech assistant professor of biology and biological engineering Rebecca Voorhees, Jason Botten of the University of Vermont, former Caltech postdoctoral scholar Devdoot Majumdar now of the University of Vermont College of Medicine, and Mitchell Guttman.
Funding was provided by the National Institutes of Health, the USC MD/PhD program, UCLA-Caltech MSTP, the American Cancer Society, the Wellcome Trust, the Office of the Vice President for Research at UVM, the Heritage Medical Research Institute, the New York Stem Cell Foundation, the Chan-Zuckerberg Initiative, and Caltech.













 0 comments
0 comments


