Credit: Courtesy of A. S. Booeshaghi
Building a map of the complex human brain and its approximately 100 billion individual neurons is no easy task. As a precursor to tackling that monumental challenge, researchers have started off with something smaller and easier—the mouse brain—in order to understand different cell types and how they are connected, and also to perfect the technological approaches to do so.
Now, a new paper describes the minute genomic details of the mouse brain at unprecedented resolution and how several types of genomics techniques were combined to enable this analysis, which was led by graduate student A. Sina Booeshaghi (MS ’19).
The study was primarily conducted in the laboratory of Lior Pachter (BS ’94), Bren Professor of Computational Biology and Computing and Mathematical Sciences, and is part of a collaboration called the Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative–Cell Census Network (BICCN), funded by the National Institutes of Health. This study and several other papers from the BICCN collaboration appear in the journal Nature on October 6.
In this new group of papers, 13 teams focused on the mouse primary motor cortex, the region of the mouse brain that controls movement.
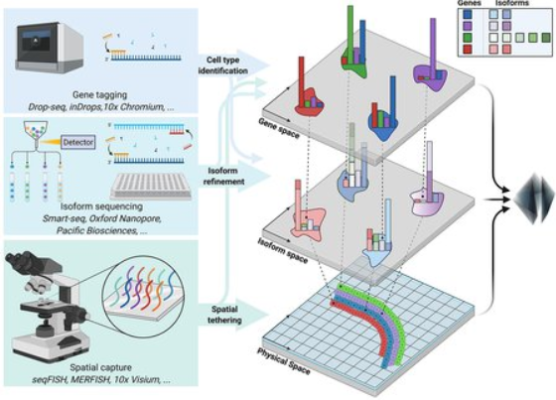
Booeshaghi’s team analyzed genomic data from brain cells collected by collaborators at the Allen Institute, a nonprofit bioscience research organization based in Seattle, Washington. By combining three different experimental techniques, each with their own strengths and weaknesses, the team was able to make detailed characterizations of gene expression in mouse cortex brain cells. The combination of techniques, Pachter says, leverages the strength of individual techniques in a way that achieves more than the sum of their parts.
The study achieved a granular level of detail by looking at what are known as gene isoforms. To understand isoforms, it is necessary to understand the expression of individual RNA transcripts that comprise genes; gene expression is the process by which the DNA of a gene is transcribed into RNA, and then RNA is read by molecular machinery to create proteins. In higher eukaryotes like humans, this flow of information includes a process called splicing, in which RNA gets chopped up and some of the pieces are glued back together before being turned into a protein. This cut-and-paste process produces many “flavors” of transcripts of the same gene: isoforms. The isoforms can translate into proteins with different functions.
By examining such isoforms, Booeshaghi and his collaborators determined that the variants are a crucial aspect contributing to functional differences between brain cells. An examination of isoforms is particularly important in the brain; the splicing process is highly active in brain tissue, and many neurological diseases result from disruptions to splicing.
The paper is titled “Isoform cell type specificity in the mouse primary motor cortex.” In addition to Booeshaghi and Pachter, additional co-authors are Zizhen Yao, Cindy van Velthoven, Kimberly Smith, Bosiljka Tasic, and Hongkui Zeng of the Allen Institute for Brain Science. Funding was provided by the National Institutes of Health Brain Initiative. Lior Pachter is an affiliated faculty member with the Tianqiao and Chrissy Chen Institute for Neuroscience at Caltech.













 0 comments
0 comments


